A Convertor from Eland2 to .WIG
Mailing
list - this is a general purpose mailing list for deep
sequencing/sequencing by synthesis.
Download - download the pacakge from our server
Bugtracker - Choose Analysis Tools | Eland2Wig
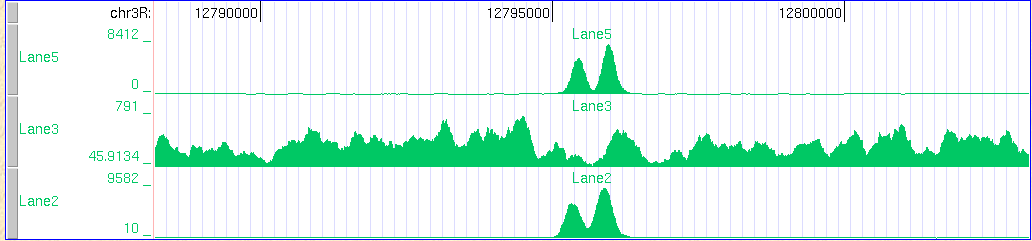
This program converts an eland 2 file to a .wig file usable at the
UCSC genome browser. The chromosome names in the eland file should be
2.fa, 3.fa etcetera. The program takes into account the forward strand
and backward strand and will from the start position on over the span
length (200 basepairs by default) increase the value in the specified
bins. The output is windowed (/binned) according to the window_size
(default = 10 bp).
Usage: eland2wig <chromosomes> <trackname> <fragmentsize> <windowsize> <strand>
The chromosome file contains the list of chromosomes that should be exported. The format is a tab seperated file. An example is given below.
2L 23011544
2LHet 368872
2R 21146708
2RHet 3288761
3L 24543557
3LHet 2555491
3R 27905053
3RHet 2517507
4 1351857
Uextra 29004656
U 10049037
X 22422827
XHet 204112
YHet 347038
The second column are the lengths of the respective chromosomes. The chromosome names in the _eland_ file should be 2.fa, 3.fa, XHet.fa etcetera
The trackname is an arbitrary name you want to give to the wiggle track.
The fragmentsize is the length of the fragments. For instance 160 bp if the retained fragments were around 200bp. (160bp because the two adaptors are around 20bp each). The fragmentsize is only use when the strand option is to take both forward and reverse alignments.
The windowsize is the size of each sample in the wigglefile. Defaults to 10, meaning to each dot in the wigglefile represents 10 nucleotides.
The strand option can be 1, 0 or -1. If it is 1 then only the forward strand will be shown, without smoothing according to the fragmentsize. If it is -1, only the reverse strand will be shown, again without fragmentsize smooting. If the strandoption is 0 then both strand will be used with an smoothed according to the fragmentsize.
For instance, to convert the output of lane 5 to a wig file use the
following:
eland2wig drosmel.genome Lane5 160 10 0 <s_5_export.txt | gzip - -9 >s_5.wig.gz
If you only want to include qualitfy filtered reads you could use something like
grep Y$ s_5_export.txt | eland2wig drosmel.genome Lane5 160 10 0 | gzip - -9 >s_5.wig.gz
Paired end reads
The output of a paired end run can also be converted to zig files. In this process we only keep the paired end reads that have matching chromosomes and for which the start and stop position are sufficiently distant (> 2x the readlength) and sufficiently small (< the required fragment length). To run such an analysis take one of the eland export files (or both of them) and pipe it through peland2wig in a similar fashion
cat s_1_1_export.txt s_1_2_export.txt | grep Y$ | peland2wig homsap_ncbi36.50.genome Lane1 1000 | gzip -9 - >s_5.wig.gz
.vec files
Some of the programs in the analysis suite require the genome signal to be saved to disk. This is done by exporting .vec files with the eland2vec tool.
Usage: eland2vec <input> [<fragmentsize>] [<strand>]
A chromosome file is not necessary for this tool. It will span the entire measured area.
For instance, to convert the output of lane 5 to a wig file use the
following:
eland2vec s_5_export.txt | gzip - >s_5.vec.gz
- http://analysis.yellowcouch.org/

